Verpeut et al. Y-maze c-Fos brain samples
This page contains links to interactive visualizations of brain-wide c-Fos expression and cerebellar injection site localization from mouse subjects studied in the manuscript “Cerebellar contributions to a brainwide network for flexible behavior” by Jessica L. Verpeut et al. (pdf: https://www.biorxiv.org/content/10.1101/2021.12.07.471685v1). Treatment groups are listed in bold, and links are for individual subjects. Only treatment groups CNO Crus I Left, CNO Crus I Right, Crus I Bilateral Reversal, and Lobule VI Reversal contain injections. In those cases, a separate “(injection)” link appears beside each c-Fos link. Any subjects with the prefix “dadult” have neither injection links nor an atlas layer (see below) due to the fact that the cerebellum was removed from the rest of the brain before imaging. c-Fos expression is still visible in those brains, however.
Neuroglancer How-to
The visualizations at the following links use the web application Neuroglancer for displaying and interacting with the data. Neuroglancer displays datasets in “layers”, each being represented by a text box near the top of the screen. The figure below shows what the top half of the screen will look like upon clicking the link an19 (c-Fos) in the CNO reversal (control) group section below.
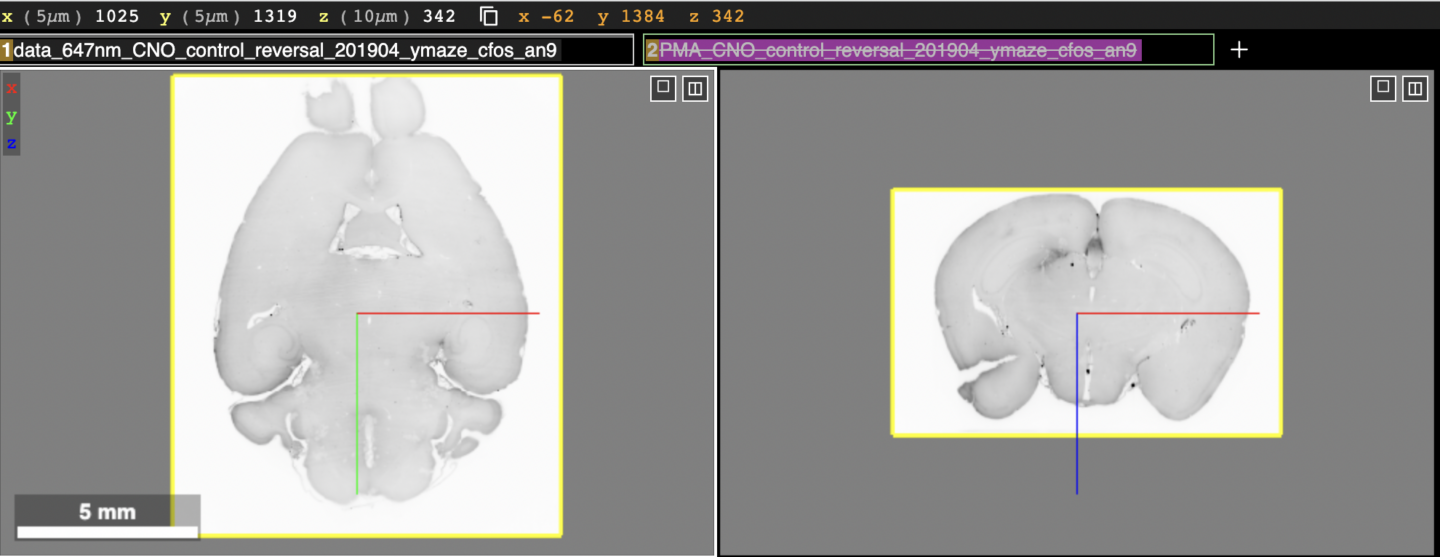
At the very top, the current position within the brain volume along with the physical scale of the dataset are shown. Below that are the layer boxes, “data_647nm_CNO_…” and “PMA_CNO_…” The first contains the brain tissue and the second the Princeton Mouse Brain Atlas (PMA) which has been aligned to the brain tissue (not shown in the figure). Below the layer boxes are the viewer panels. In the figure above the horizontal (left) and coronal (right) sections of the same brain tissue are shown. The sagittal view (not shown in the figure) is displayed in a lower right panel in Neuroglancer. The 3D view, which can be ignored for this dataset (not shown in the figure) is displayed in the lower left panel in Neuroglancer. There is also an x,y,z coordinate axis (blue, red, green lines) that indicates the center of both views. It can be used to determine the relative orientation between the two views.
In the above figure, the brain atlas layer is toggled off by default, indicated by the strikethrough text in its layer box. To toggle on the brain atlas, click the text box where the strikethrough text is shown. Upon doing that, the top half of the screen will now look like this:
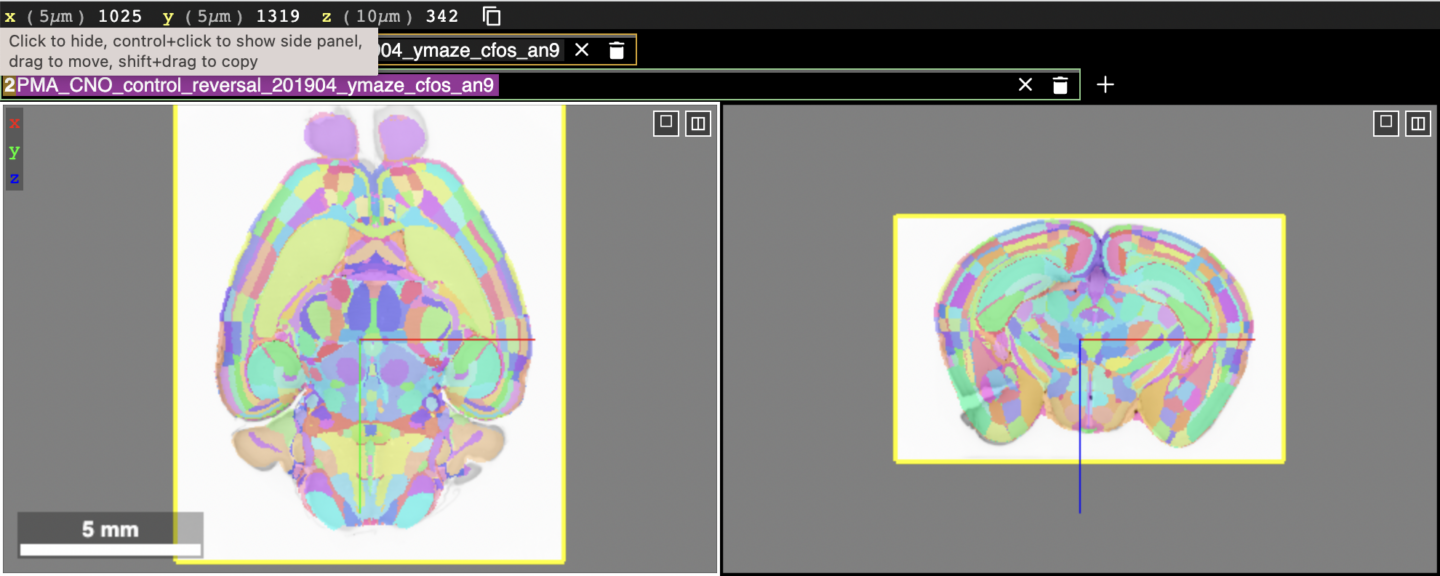
The aligned brain atlas regions are now shown on top of the brain tissue. They are multicolored so that the boundaries between regions can be distinguished. Hovering over a brain region will display the name of the region in the atlas in the layer text box for the atlas. The atlas layer can be made more transparent so that the brain tissue underneath can be seen more easily. This can be done by selecting the “Render” tab on the right hand side panel (not shown in figure but will be displayed in Neuroglancer) and adjusting the opacity slider.
The injection links below have a third layer, which shows the segmented area from the injection site we detected using a program for analysis. The following figure shows what those links will look like:
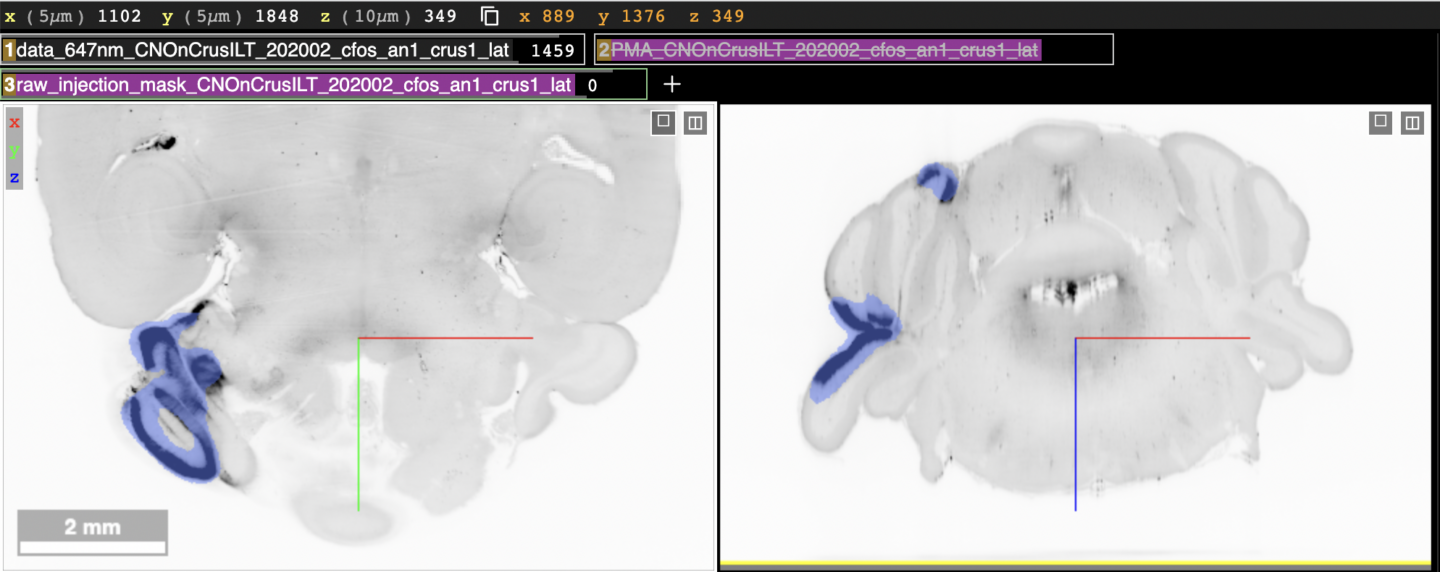
The first two layers “data_647nm_…” and “PMA_CNO…” have the same layout as before (but for a different subject), where the brain atlas layer (second layer) is toggled off by default. The third layer, “raw_injection_mask_…” is new. This layer is shown as the blue area in the viewer panels.
Neuroglancer has some basic keystroke commands that enable you to explore the multidimensional data:
- h key: open help menu
- (2D view) left-click and drag: translate section
- (2D and 3D view) mouse-wheel: scroll along anterior-posterior (AP) axis. Trackpad: two finger scroll
- (2D and 3D view) shift+mouse-wheel: scroll (x10 speed) along anterior-posterior (AP) axis.
- (2D and 3D view) ctrl+mouse-wheel or ctrl+”+/-“: zoom in and out
- (2D and 3D view) double left-click: select/de-select segment
- (2D and 3D view) Hover over a segment: highlights the region and displays the brain region name in the atlas layer box area.
- (3D view) left-click and drag: rotate brain
- (3D view) s key: toggle the gray reference slice on/off
- right click on layer box: open up right side panel of options for displaying that layer such as opacity, color scheme, segment selection
CNO reversal (control)
an9 (c-Fos)an10 (c-Fos)
an11 (c-Fos)
an12 (c-Fos)
an13 (c-Fos)
an14 (c-Fos)
an15 (c-Fos)
an16 (c-Fos)
an17 (c-Fos)
an18 (c-Fos)
CNO Crus I Left
an1 (c-Fos) (injection)an2 (c-Fos) (injection)
an5 (c-Fos) (injection)
an6 (c-Fos) (injection)
an7 (c-Fos) (injection)
an8 (c-Fos) (injection)
an9 (c-Fos) (injection)
an15 (c-Fos) (injection)
an16 (c-Fos) (injection)
an17 (c-Fos) (injection)
CNO Crus I Right
an3 (c-Fos) (injection)an4 (c-Fos) (injection)
an10 (c-Fos) (injection)
an11 (c-Fos) (injection)
an12 (c-Fos) (injection)
an13 (c-Fos) (injection)
an14 (c-Fos) (injection)
an18 (c-Fos) (injection)
an19 (c-Fos) (injection)
an20 (c-Fos) (injection)
Acquisition day 1
an011 (c-Fos)an012 (c-Fos)
an013 (c-Fos)
an014 (c-Fos)
an015 (c-Fos)
an016 (c-Fos)
an017 (c-Fos)
an018 (c-Fos)
an019 (c-Fos)
an020 (c-Fos)
Crus I Bilateral Reversal
092220_vectorandcrusIbil_openfieldonly-001 (c-Fos) (injection)092220_vectorandcrusIbil_openfieldonly-002 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-003 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-004 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-005 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-005 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-006 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-007 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-008 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-009 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-010 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-011 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-012 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-013 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-014 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-015 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-016 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-017 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-018 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-019 (c-Fos) (injection)
092220_vectorandcrusIbil_openfieldonly-020 (c-Fos) (injection)
dadult_pc_crus1_2
dadult_pc_crus1_5
dadult_pc_crus1_6
dadult_pc_crus1_7
dadult_pc_crus1_8
dadult_pc_crus1_9
Habituation
an001 (c-Fos)an002 (c-Fos)
an003 (c-Fos)
an004 (c-Fos)
an005 (c-Fos)
an006 (c-Fos)
an007 (c-Fos)
an008 (c-Fos)
an009 (c-Fos)
an010 (c-Fos)
Lobule VI Reversal
an19 (c-Fos) (injection)an21 (c-Fos) (injection)
an22 (c-Fos) (injection)
an23 (injection)
an24 (c-Fos) (injection)
an25 (c-Fos) (injection)
an26 (No injection site availabile)
an27 (c-Fos) (injection)
an28 (c-Fos) (injection)
dadult_pc_lob6_13
dadult_pc_lob6_15
dadult_pc_lob6_17
dadult_pc_lob6_18
dadult_pc_lob6_19
dadult_pc_lob6_20
dadult_pc_lob6_21